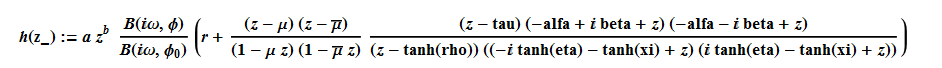
(1)
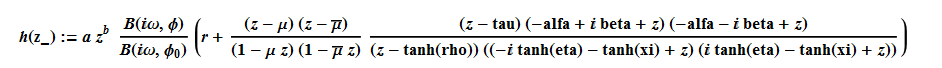 |
(1) |
|
cx-scale×Bessel-f. × (mix +
all-pass
×
poles-and-zeros
) |
21 ^ o-110910/zadm-0.01745-z00.sp input complex admittance and its confidenceOutput:
22 ^ o-110910/cohc-0.01745-z00.sp input coherence
23 ^ o-110910/zpha-0.01745-z00.sp input confidence of phase
31 < o-110910/zadm-0.01745-z00-ra.sp output
81 < fitpzm.txt debug output if qdbg_polezero = .true.
q
splist o-110910/zadm-0.01745-z00-ap.sp xCAD : The data after tshift and qconjgsp operations
<OPENF->3> 21:^ o-110910/zadm-0.01745-z00-ap.sp -
<getsp->d> Header: 360 CAD gravseis T 3.178211E-03 629.285 2.777777777777778E-004 0.000000000000000 0.000000000000000
<GetsSt>>> Unit 21 tag CAD. VarRedFac= 0.0032, DegFreed= 629.28
<GetsCn>>> conf.limits: 0.0000E+00 0.0000E+00
<Main-->>> read done, n= 360 dt= 2.7778E-04 stats: T 3.1782E-03 6.2928E+02 conf: 0.0000E+00 0.0000E+00
<GETSP->>> CMPX #21:o-110910/zadm-0.01745-z00-ap.sp - contains ->CAD<>gravseis>< <- bins 0... 360
<GETSP->>> CMPX #21:o-110910/zadm-0.01745-z00-ap.sp - contains ->CAM<>gravseis>< <- bins 0... 360
<GETSP->>> CMPX #21:o-110910/zadm-0.01745-z00-ap.sp - contains ->CAR<>gravseis>< <- bins 0... 360
<GETSP->>> REAL #21:o-110910/zadm-0.01745-z00-ap.sp - contains ->ADC<>gravseis>< <- bins 0... 360
<GETSP->>> REAL #21:o-110910/zadm-0.01745-z00-ap.sp - contains ->APH<>gravseis>< <- bins 0... 360
<GETSP->>> EOF #21:o-110910/zadm-0.01745-z00-ap.sp
____________________________________________________________________________
Parameter - type - explanation [default]
____________________________________________________________________________
Scaling and mixing:
zamps(50) - c16 - Re(zamp(i)) => a , Im(zamp(i)) => b [50*(1.0,0.0)]
r(50) - r8 - r(i) => r [50*1.0]
Allpass:
zap - c16 - (amp,pha[deg]) of starting value for μ in (1) [√2,45]
Bessel filter:
bdf0 - r8 - the design parameter used in data preparation [0.01745]
bdfa - r8 - the starting value for the adjustment [0.01745]
Up to 50 pole and zero parameters of each kind,
specifying the starting values for iteration:
xi(50) - r8 - complex pole, real part [50*0.5]
eta(50) - r8 - complex pole, imag part [50*0.5]
rho(50) - r8 - real pole [none]
alfa(50) - r8 - complex zero, real part [none]
beta(50) - r8 - complex zero, imag part [none]
tau(50) - r8 - real zero [none]
Extension of equation (1), poles and zeros:
nextend - i - how many successive expansions of (1)
For each i from 1 to nextend:
mallp(50) - i - ... (1 or 0) if an all-pass is to be adjusted
or not [50*0]
mbs[50] - i - ... (1 or 0) if the Bessel filter (ratio of
B()'s in (1)) is to be adjusted or not [50*0]
ma(50) - i - ... ma(i)=0 => r in (1) is fixed,
... ma(i)=1 => r in (1) is relaxed [50*1]
mb(50) - i - ... (1 or 0) if b is to be solved or not [50*1]
mp(50) - i - ... how many complex poles [50*0]
mrp(50) - i - ... real poles [50*0]
mz(50) - i - ... complex zeros [50*0]
mrz(50) - i - ... real zeros [50*0]
Files and data:
tshift - r8 - incur the effect of a time shift on the input
spectrum, in seconds [0.0d0]
qconjgsp - l - transpose the spectrum (cx-conjug) [.false.]
iu_csp - i - file unit of complex admittance spectrum [21]
iu_cohc - i - ... of coherence spectrum [21]
iu_pha - i - ... of phase spectrum (needed to copy uncertainty) [23]
iu_out - i - file unit for output [31]
tagc - s3 - id-tag of complex admittance ['CAD']
tagh - s3 - ... coherence ['KOH']
taga - s3 - ... confidence of admittance (to copy to output) ['ADC']
tagp - s3 - ... confidence of phase (to copy to output) ['APH']
Iteration control:
nfcohc - i - Data weighting:
n0cohc - i set coherence=1 from 0 to nfcohc-1
and from n0cohc to the end [0,ndim]
iter(50) - i - maximum number of iterations in Numerical
Recipes subroutine dfpmin [50*50]
iter(i)=0 will bypass iteration; useful for
forward computation
gtol - r8 - convergence control for dfpmin [1.d-6]
Print and debug:
msglvl - i - 0 will print all debug messages due to getsp/outsp
4 will print none [4]
qpr_parput - l - print the parameters p() at every stage of
extension [.false.]
qdbg_parset - l - debug option in polezero subroutine during
model extension.
Output to unit 81 as under qdbg_polezero
is only preserved if qdbg_polezero=.false. [.false.]
qdbg_polezero - l - (subject to modification - check zpolezero
etc. in pzfit.f, qdbg, for what is in effect).
Idea: the different terms of (1) in final form
are printed to file unit 81 [.false.]
qdbg_iter(50) - l - print like qdbg_polezero during iteration
of extension stage i if qdbg_iter(i)=.true. [.false.]
qdbg_fpmin(50)- l - debug option in Numerical Recipes subroutine
dfpmin in extension step i [50*.false.]
qafter_math - l - vary a complex parameter in a 2-d loop (radius and angle)
or a real parameter between -1 and +1,
and output the misfit (10log(chi2'-chi2) to
file unit 62. The last added zero or pole can
be varied, alternately the all-pass parameter:
qam_allpass - l -
qam_pole - l -
qam_zero - l -
qam_rpole - l -
qam_rzero - l -
____________________________________________________________________________
21 ^ o-110910/zadm-0.01745-z00.spAfter-math
22 ^ o-110910/cohc-0.01745-z00.sp
23 ^ o-110910/zpha-0.01745-z00.sp
31 < o-110910/zadm-0.01745-z00-ra.sp
q
¶m
iu_cohc=22
msglvl=3
qdbg_polezero=.true.
qdbg_fpmin=3*.false.
nextend=3
mp=0,1,2
mb=0,1,1
ma=0,1,1
r=0.0d0,1.0d0
zamps=(1.0d0,0.0d0),(1.0d0,0.01d0),(-3.0082E-01,-3.4378E-02)
&end
splist-plot -o -X0,1 -gain -P ~/www/4me/cal-seis$SUBDIR -O tmp -E \
+ -Hz -f0 -adB \
++ o-110910/zadm-0.01745-z00-ra.sp CAM ADC
or
plot-fitpz-result ra